Research
The Zheng Lab uses a wide range of tools and systems in their work, including genetics in model organisms, model organism development, cell culture, biochemistry, proteomics, and genomics. In recent years, their findings have broadened their research scope to include three research areas: 1) The mechanism of cell division. 2) The mechanism of genome organization in development, homeostasis, and aging. 3) The mechanism of endosymbiosis in cnidaria.
Current Topics
The way a progenitor cell partitions itself during cell division has a profound influence on the behavior and fate of its daughter cells. Understanding this partitioning requires us to study both the mechanism of equal chromosome segregation and the means a dividing cell segregates critical cell fate determinants into daughter cells. The mitotic spindle apparatus is one of the most complex cellular machines consisting of microtubules, microtubule-associated proteins (MAPs), and motors. The spindle also associates with many poorly defined proteins and membranes. Historically these spindle-associated materials are called the spindle matrix. The importance of the spindle matrix and the value of studying it have remained a subject of debate.
We have uncovered protein complexes called γ-tubulin ring complex (γTuRC) and γ-tubulin small complex (γTuSC) that mediate microtubule nucleation and organization in mitotic and interphase cells. Through the study of microtubule nucleation, we became fascinated by the more complex and dynamic behaviors of microtubules during mitotic spindle assembly. By using the powerful Xenopus egg extract, we and others have uncovered an important signaling pathway mediated by the nuclear small GTPase Ran that regulates multiple aspects of cell division. We show that RanGTPase also regulates the assembly of the spindle matrix containing lamin-B. Based on our studies, we propose that RanGTP and the spindle matrix promote both spindle assembly and orientation. Consistent with this, we show that the spindle matrix component lamin-B regulates spindle orientation in neural stem cells in the developing mouse brain. Lamin-B may do so in part by regulating centrosome positioning.
The complexity of the spindle matrix has made the study of its structure and function relationship very difficult, which contributes to the debate of its function and even existence. By studying another spindle matrix component BuGZ, which we discovered through proteomic analyses of the Xenopus spindle matrix, we show that protein phase separation/transition represents a biophysical property of the spindle matrix. The phase separation of BuGZ along spindle microtubules promotes spindle matrix assembly, which in turn facilitates spindle microtubule assembly by concentrating tubulin. This finding opens the door to further characterize the structure and function of the spindle matrix in cell division.
The nuclear lamina and chromatin-bound proteins are known to regulate genome organization in interphase cells, yet how cells in different lineages acquire and maintain their unique genome architecture has remained poorly understood. We use various tools in genetics, genomics (such as ChIP-seq, RNA-seq, single cell RNA-seq, and Hi-C), cell biology, and biochemistry to study how genomes obtain their organization in stem cells (including ES cells) and differentiated cells isolated from tissues. We also analyze whether such organization plays a role in lineage specification or terminal differentiation, how such organization is maintained in adulthood, and whether genome dis-organization leads to age-associated diseases. For example, our recent studies demonstrate that lamin-B (the major structural component of the nuclear lamina) is not required for early lineage specification during development, but it is essential for proper organogenesis. Aging-associated lamin-B reduction in Drosophila fat bodies (equivalent to human fat and liver) leads to system inflammation and gut hyperplasia. These and other published and ongoing studies in the lab are allowing us to dissect the role of genome organization in the context of development, tissue function, and aging.
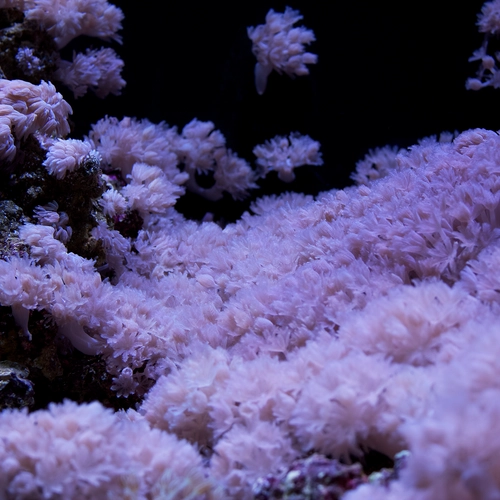
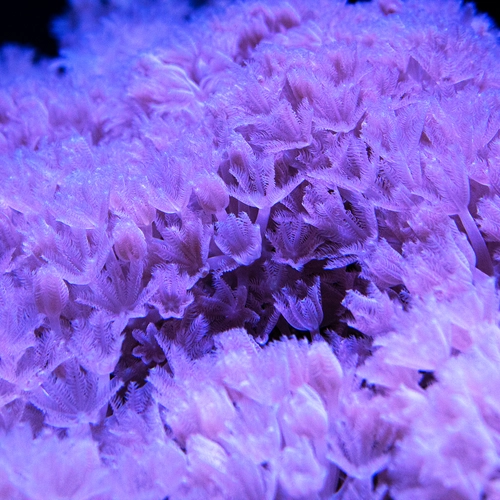
Many cnidaria species, including Hydra, upside-down jellyfish, and hard and soft corals harbor algae for photosynthesis. The algae live inside coral cells in a specialized membrane compartment called symbiosome, which shares the photosynthetically fixed carbon with coral host cells, while host cells provide inorganic carbon for photosynthesis. The molecular pathways in cnidaria cells that orchestrate algal recognition, uptake, nutrient sharing, and maintenance remain poorly understood. We have built facilities to grow various cnidaria species and have begun to create model organisms to understand cnidaria endosymbiosis.
Timeline
CV
- 1980 – 1984 | B.S. in Genetics, Sichuan University, Sichuan, China.
- 1987 – 1992 | Ph.D. in Molecular Genetics, The Ohio State University, Columbus, OH (Advisor: Berl Oakley).
- 1992 – 1996 | Postdoctoral Fellow, University of California, San Francisco, CA (Advisors: Bruce Alberts and Tim Mitchison).
- 1984 – 1986 | Lecturer, Southwestern Agricultural University, Sichuan, China.
- 1996 – Present | Staff Member/Investigator, Department of Embryology, Carnegie Institution for Science, Baltimore, MD.
- 1996 – Present | Adjunct Assistant Professor, Associate Professor (2002), Professor (2007), Department of Biology, Johns Hopkins University.
- 2000 – 2012 | Adjunct Assistant Professor, Associate Professor (2002), Professor (2007), Department of Molecular Biology and Genetics, Johns Hopkins University School of Medicine.
- 2000 – 2012 | HHMI Investigator, Howard Hughes Medical Institute.
- 2016 – 2017 | Acting Director, Dept. of Embryology, Carnegie Institution for Science.
- 2017 – Present | Director, Dept. of Embryology, Carnegie Institution for Science.
- 2018 – 2018 | Interim Co-President, Carnegie Institution for Science.
- 1997 – 2001 | PEW Scholar Award (Awarded by PEW Charitable Trusts).
- 1997 – Present | NIH R01 and R21 Research Grants.
- 1999 | Women in Cell Biology Award (Junior) (Awarded by the American Society for Cell Biology).
- 2000 – 2012 | HHMI Investigator, Howard Hughes Medical Institute.
- 10/2006 | Douglas D. McGregor Research Keynote Speaker, Cornell University
- 07/2007 | Keynote Speaker, Gordon Conference - Motile & Contractile Systems.
- 2008 | National Associate of the National Research Council, the National Academies of Sciences (An honorary lifetime appointment for extraordinary service to the National Research Council of the National Academies of Sciences).
- 2012 – 2016 | Senior Scholar in Aging, Ellison Medical Foundation.
- 2017 | Fellow, American Society for Cell Biology (ASCB).
- 2020 – 2025 | Investigator, Gordon and Betty Moore Foundation (Symbiosis in Aquatic Systems).
- 2022 – Present | Member, R35 awards to early career investigators, Center for Scientific Review, NIH.
- 2013 – 2020 | Member, NCSD Study Section (Nuclear and Cytoplasmic Structure/Function and Dynamics), Center for Scientific Review, NIH.
- 2016 – 2019 | Chair, International Affairs Committee, American Society of Cell Biology.
- 2018 – 2021 | Member, Wellcome Trust Interview Panel (for Grant Funding).
- 2013 – Present | National Scientific Advisory Council (NSAC); American Federation for Aging Research (AFAR).
- 2011 – 2013 | Council Member, the Council of American Society for Cell Biology (ASCB).
- 2011 – 2013 | Member, International Affairs Committee, American Society for Cell Biology.
- 2007 – 2008 | Member, Program Committee, American Society for Cell Biology.
- 2005 – 2007 | Study Section NDT (Nuclear Dynamics and Transport), Center for Scientific Review (CSR), NIH.
- 2003 – 2004 | Study Section CDF4 (Cell Development and Function 4), Center for Scientific Review (CSR), NIH.
- 1999 – 2006 | Member, Woman in Cell Biology Committee, American Society for Cell Biology.
- 2001 – 2004 | Member of Organizing Committee, The Chinese-American Frontiers of Science, a special program of the US National Academy of Sciences.
- 2000 – 2002 | Prostate Cancer CBY2 Review Group, Department of Defense, United States.
- 2002 | Review Committee for Northwestern University School of Medicine
- Department of Cell Biology.
- 2001 | Review committee for International Scholar’s Program, Howard Hughes Medical Institute.
- 2001 | Chair, Local Arrangements Committee, 41st Annual Meeting, The American Society for Cell Biology, December 8-12, 2001, DC.
Recent Publications
All Publications
1. Zheng Y, Jung MK, & Oakley BR (1991). g-tubulin is present in Drosophila melanogaster and Homo sapiens and is associated with the centrosome. Cell 65:817-823.
2. Zheng Y, Wong ML, Alberts B, & Mitchison TJ (1995). A g-tubulin ring complex from the unfertilized egg of Xenopus laevis can nucleate microtubule assembly in vitro. Nature 378:578-583. (News & Views: Oakley, Nature 378:555-556)
3. Wilson PG, Zheng Y, Oakley CE, Oakley BR, Borisy GG, & Fuller MT (1997). Differential expression of two g-tubulin isoforms during gametogenesis and development in Drosophila. Developmental Biology 184:207-221.
4. Dictenberg JB, Zimmerman W, Sparks CA, Young A, Vidair C, Zheng Y, Carrington W, Fay FS, & Doxsey SJ (1998). Pericentrin and g-Tubulin Form a Protein Complex and Are Organized into A Novel Lattice at the Centrosome. Journal Cell Biology 141:163-174.
5. Martin O, Gunawardane R., Iwamatsu A, & Zheng Y (1998). Xgrip109: A g-tubulin associated protein with an essential role in gTuRC assembly and centrosome function. Journal of Cell Biology 141:675-687.
6. Moritz M, Zheng Y, Alberts B, & Oegema K (1998). Recruitment of the g-tubulin ring complex to Drosophila salt-stripped centrosome scaffolds. Journal of Cell Biology 142:775-786.
7. Zheng Y, Wong ML, Alberts B, & Mitchison T (1998). Purification and assay of g-tubulin ring complex. Methods in Enzymology 298, Part B, 218-228.
8. Field CM, Oegema K., Zheng Y, Mitchison T, & Walczak CE (1998). Purification of Cytoskeletal Proteins Using Peptide Antibodies. Methods in Enzymology 298, Part B, 525-541.
9. Oegema K, Wiese C, Martin OC, Milligan RA, Iwamatsu A, Mitchison T, & Zheng Y (1999). Characterization of Two Related Drosophila g-tubulin Complexes that Differ in Their Ability to Nucleate Microtubules. Journal of Cell Biology 144:721-733.
10. Wiese C & Zheng Y (1999). g-Tubulin Complexes and Their Interaction with Microtubule Organizing Centers. Current Opinion in Structural Biology 9:250-259.
11. Wilde A & Zheng Y (1999). Stimulation of Microtubule Aster Formation and Spindle Assembly in Xenopus Egg Extracts by the Small GTPase Ran. Science 284:1359-1362. (News Focus: Pennisi, Science 284:1260-1261, 1999; Commentary: Desai & Hyman, Current Biology 9:R704-707, 1999)
12. Wiese C & Zheng Y (2000). A New Function for the g-tubulin Ring Complex as a Microtubule Minus-end Cap. Nature Cell Biology 2:358-364. (News & Views: Erickson, Nature Cell Biology 2:E93-E96)
13. Zhang L, Keating T, Wilde, A, Borisy G, & Zheng Y (2000). The Role of Xgrip210 in g-Tubulin Ring Complex Assembly and Centrosome Recruitment. Journal of Cell Biology 151:1525–1535.
14. Gunawardane R, Martin O, Cao K, Zhang L, Dej K, Iwamatsu A, & Zheng Y (2000). Characterization and Reconstitution of Drosophila g-Tubulin Ring Complex Subunits. Journal of Cell Biology 151:1513–1523.
15. Gunawardane RN, Lizarraga SB, Wiese C, Wilde A, & Zheng Y (2000). g-Tubulin Complexes and Their Role in Microtubule Nucleation. Current Topics in Developmental Biology 49:55-73.
16. Wilde A, Lizarraga S, Zhang L, Wiese C, Gliksman N, Walczak C, & Zheng Y (2001). Ran stimulates spindle assembly by changing microtubule dynamics and the balance of motor activities. Nature Cell Biology 3:221-227. (News & Views: Walczak, Nature Cell Biology 3:E69-70, 2001)
17. Wiese C, Wilde A, Adam S, Moore M, Merdes A, & Zheng Y (2001). Role of Importin-b in Coupling Ran to Downstream Targets in Microtubule Assembly. Science 291:653-656. (News & Views: Walczak, Nature Cell Biology 3:E69-70, 2001)
18. Gunawardane RN, Zheng Y, Oegema K, & Wiese C (2001). Purification and reconstitution of Drosophila gamma-tubulin complexes. Methods in Cell Biology 67:1-25.
19. Lizarraga SB, Zheng Y, & Wilde AR (2002). Characterization of the effects of RanGTP on the microtubule cytoskeleton. Methods in Molecular Biology 189:247-260.
20. Gunawardane R, Martin OC, & Zheng Y (2003). Characterization of a new gTuRC subunit with WD repeats. Molecular Biology of the Cell 14:1017-1026.
21. Tsai MY, Wiese C, Cao K, Martin OC, Donovan P, Ruderman J, Prigent C, & Zheng Y (2003). A Ran-signaling pathway mediated by the mitotic kinase Aurora A in spindle assembly. Nature Cell Biology 5:242-248.
22. Li HY, Wirtz D, & Zheng Y (2003). A mechanism of coupling RCC1 mobility to RanGTP production on the chromatin in vivo. Journal of Cell Biology 160:635-644.
23. Li HY, Cao K, & Zheng Y (2003). Ran in spindle checkpoint: a new function for a versatile GTPase. Trends in Cell Biology 13:553-557.
24. Cao K, Nakajima R, Meyer HH, & Zheng Y (2003). The AAA-ATPase Cdc48/p97 regulates spindle disassembly at the end of mitosis. Cell 115:355-367. (Highlight: Nature Reviews Molecular Cell Biology 4:906, 2003; Commentary: Cheeseman and Desai, Current Biology 14:R70-72, 2004)
25. Ems-McClung SC, Zheng Y, & Walczak CE (2004). Importin / and Ran-GTP Regulate XCTK2 Microtubule Binding through a Bipartite Nuclear Localization Signal. Molecular Biology of the Cell 15:46-57.
26. Kawaguchi S & Zheng Y (2004). Characterization of a Drosophila Centrosome Protein CP309 That Shares Homology with Kendrin and CG-NAP. Molecular Biology of the Cell 15:37-45.
27. Li HY & Zheng Y (2004). Phosphorylation of RCC1 in mitosis is essential for RanGTP gradient production and spindle assembly in mammalian cells. Genes and Development 18:512-527.
28. Cao K & Zheng Y (2004). The Cdc48/p97-Ufd1-Npl4 Complex: Its Potential Role in Coordinating Cellular Morphogenesis during the M-G1 Transition. Cell Cycle 3:422-424.
29. Li HY & Zheng Y (2004). The Production and Localization of GTP-Bound Ran in Mitotic Mammalian Tissue Culture Cells. Cell Cycle 3:993-995.
30. Ducat DC & Zheng Y (2004). Aurora kinases in spindle assembly and chromosome segregation. Experimental Cell Research 301:60-67.
31. Nakajima R, Tsai M-Y, & Zheng Y (2004). Centrosomes and Microtubule Nucleation. Encyclopedia of Biological Chemistry 1:372-376. W. J. Lennarz and M. D. Lane (Ed), Elsevier Inc. (not listed in PubMed)
32. Zheng Y (2004). G Protein Control of Microtubule Assembly. Annual Review of Cell and Developmental Biology 20:867-894.
33. Tsai M-Y & Zheng Y (2005). Aurora A Kinase-Coated Beads Function as Microtubule-Organizing Centers and Enhance RanGTP-Induced Spindle Assembly. Current Biology 15:2156-2163. (Highlighted in Meeting Report: Nuclear protein supports spindle. Journal of Cell Biology 172:488, 2006)
34. Vong QP, Cao K, Li HY, Iglesias PA, & Zheng Y (2005). Chromosome Alignment and Segregation Regulated by Ubiquitination of Survivin. Science 310:1499-1504. (Perspective: Earnshaw, Science 310:1443-1444)
35. Tsai M-Y, Wang S, Heidinger JM, Shumaker D, Adam SA, Goldman RD, & Zheng Y (2006). A Mitotic Lamin B Matrix Induced by RanGTP Required for Spindle Assembly. Science 311:1887-1893. (Lead article; News & Views: Hayes, Nature Cell Biology 8:550, 2006; Research Highlight: Nature Reviews Molecular Cell Biology 7:307, 2006).
36. Goodman B & Zheng Y (2006). Mitotic spindle morphogenesis: Ran on the microtubule cytoskeleton and beyond. Biochemical Society Transactions 34:716-721.
37. Wiese C & Zheng Y (2006). Microtubule nucleation: g-tubulin and beyond. Journal of Cell Science 119:4143-4153.
38. Zheng Y & Tsai M-Y (2006). The Mitotic Spindle Matrix: A Fibro-Membranous Lamin Connection. Cell Cycle 5:2345-2347.
39. Spradling AC & Zheng Y (2007). The Mother of All Stem Cells? Science 315:469-470.
40. Liu Z, Vong QP, & Zheng Y (2007). CLASPing Microtubules at the trans-Golgi Network. Developmental Cell 12:839-840.
41. Li HY, Ng WP, Wong CH, Iglesias PA, & Zheng Y (2007). Coordination of Chromosome Alignment and Mitotic Progression by the Chromosome-Based Ran Signal. Cell Cycle 6:1886-1895.
42. Channels WE, Nedelec FJ, Zheng Y, & Iglesias PA (2008). Spatial regulation improves anti-parallel microtubule overlap during mitotic spindle assembly. Biophys Journal 94:2598-2609.
43. Zheng Y & Oegema K (2008). Cell Structure and Dynamics. Current Opinion in Cell Biology 20:1–3. Editor (not listed in PubMed)
44. Ducat D, Kawaguchi S, Liu H, Yates JR 3rd, & Zheng Y (2008). Regulation of microtubule assembly and organization in mitosis by the AAA+ ATPase Pontin. Molecular Biology of the Cell 19:3097-3110.
45. Wilde A & Zheng Y (2009). Ran out of the nucleus for apoptosis. Nature Cell Biology 11:11-12.
46. Li M, Tsai MY, Lu B, Chen R, Yates III JR, Zhu X, & Zheng Y (2009). A Requirement of Nudel and Dynein for Spindle Matrix Assembly during Spindle Morphogenesis. Nature Cell Biology 11:247-256.
47. Martin O, DeSevo CG, Guo BZ, Koshland DE, Dunham MJ, & Zheng Y (2009). Telomere behavior in a hybrid yeast. Cell Research 19:910-912.
48. Liu Z & Zheng Y (2009). A Requirement for Epsin in Mitotic Membrane and Spindle Organization. Journal of Cell Biology 186:473-480.
49. Bembenek JN, White JG, & Zheng Y (2010). A Role for Separase in the Regulation of RAB-11-positive Vesicles at the Cleavage Furrow and Midbody. Current Biology 20:259-264. (Commentary: Lopez-Aviles S and Uhlmann F, The Art of Multi-Tasking, Current Biology 20:R101-103, 2010)
50. Wallingford JB, Liu KJ, & Zheng Y (2010). Xenopus. Current Biology 20:R263-4.
51. Vong QV, Liu Z, Yoo JG, Chen R, Xie W, Sharov AA, Fan CM, Liu C, Ko MSH, & Zheng Y (2010). A Role for Borg5 during Trophectoderm Differentiation. Stem Cells 28:1030-1038.
52. Zheng Y (2010). Mitotic spindle matrix may hold the answer to orchestrating cell division. Nature Reviews Molecular Cell Biology 11:529-535.
53. Goodman B, Channels W, Qiu M, Iglesias P, Yang G, & Zheng Y (2010). Lamin-B3 counteracts the kinesin Eg5 to restrain spindle pole separation during spindle assembly. J Biol Chem 285:35238-44.
54. Poirier CC, Zheng Y, & Iglesias PA (2010). Biophysical J. Mitotic Membrane Helps to Focus and Stabilize the Mitotic Spindle. Biophys J 99:3182-3190.
55. Hoang ML, Tan FJ, Lai DC, Celniker SE, Hoskins RA, Dunham MJ, Zheng Y, Koshland D (2010). Competitive Repair by Naturally Dispersed Repetitive DNA during Non-Allelic Homologous Recombination. PLoS Genet 6(12):e1001228.
56. Wang S & Zheng Y (2011). Identification of a novel dynein-binding domain in Nudel essential for spindle pole organization in Xenopus egg extracts. J Biol Chem 286:587-93.
57. McDole K, Xiong Y, Iglesias PA, Zheng Y (2011). Lineage mapping the pre-implantation mouse embryo by two-photon microscopy, new insights into the segregation of cell fates. Dev Biol. 355:239-49.
58. Kang J, Goodman B, Zheng Y, & Tantin D (2011). Dynamic Regulation of Oct1 during Mitosis by Phosphorylatino and Ubiquitination. PloS One 6 (8), e23872.
59. Kim Y, Sharov AA, McDole K, Cheng M, Hao H, Fan C-M, Gaiano N, Ko MSH, & Zheng Y (2011). Mouse B-type lamins are required for proper organogenesis but not by ES cells. Science 334:1706-10. (Commentary: David R, “Nucleoskeleton - Uncovering roles for lamin B,” Nat Rev Mol Cell Biol 13(1):3, 2011)
60. Kim Y, McDole K, and Zheng Y (2012). The function of lamins in the context of tissue building and maintenance. Nucleus 3:256-62.
61. McDole K and Zheng Y (2012). Generation and live imaging of an endogenous Cdx2 reporter mouse line. Genesis 50:775-782.
62. Liu Z, Vong QP, & Zheng Y (2012). Using ES Cells Labeled with GFP for Analyzing Cell Behavior During Differentiation. Curr Protoc Stem Cell Biol. Chapter 1:Unit1D.8.
63. Jia J, Zheng X, Hu G, Cui K, Zhang A, Zhang J, Du Y, Liu C, Zhao K, and Zheng Y (2012). Regulation of pluripotency and self-renewal of ES cells through epigenetic threshold modulation and mRNA pruning. Cell 151:576–589. (Commentary: Laskowski AI & Knoepfler PS, “Utf1: Goldilocks for ESC bivalency,” Cell Stem Cell 11:732-734, 2012; Muers M, “Gene regulation: bivalency buffer makes pluripotency connections.” Nat Rev Genet 13:826-827, 2012)
64. Zheng Y & Iglesias PA (2013). Nucleating new branches from old. Cell 152:669-670.
65. Chen H, Chen X, and Zheng Y (2013). The nuclear lamina regulates germline stem cell niche organization via modulation of EGFR signaling. Cell Stem Cell 13:73-86.
66. Wang S, Ketcham SA, Schön A, Goodman B, Wang Y, Yates J 3rd, Freire E, Schroer TA, Zheng Y (2013). Nudel/NudE and Lis1 promote dynein and dynactin interaction in the context of spindle morphogenesis. Mol Biol Cell 24:3522-3533.
67. Kim Y, Zheng X, & Zheng Y (2013). Proliferation and differentiation of mouse embryonic stem cells lacking all lamins. Cell Res 23:1420-1423.
68. Kim Y & Zheng Y (2013). Generation and characterization of a conditional deletion allele for Lmna in mice. Biochem Biophys Res Commun 440:8-13.
69. Guo Y & Zheng Y (2013). Sculpting the nucleus with dynamic microtubules. Dev Cell 27:1-2.
70. Liu Z, Vong QP, Liu C, & Zheng Y (2014). Borg5 is required for angiogenesis by regulating persistent directional migration of the cardiac microvascular endothelial cells. Mol Biol Cell 25:841-51.
71. Jiang H, He X, Wang S, Jia J, Wan Y, Wang Y, Zeng R, Yates III J, Zhu X, & Zheng Y (2014). A Microtubule-Associated Zinc Finger Protein, BuGZ, Regulates Mitotic Chromosome Alignment by Ensuring Bub3 Stability and Kinetochore Targeting. Dev Cell 28:268-281.
72. Chen Y and Zheng Y (2014). Nuclear lamina builds tissues from the stem cell niche. Fly 8:63-67.
73. Guo Y, Kim Y, Shimi T, Goldman RD, Zheng Y (2014). Concentration-dependent lamin assembly and its roles in the localization of other nuclear proteins. Mol Biol Cell, 25:1287-1297.
74. Shi C, Channels WE, Zheng Y, and Iglesias PA (2014). A computational model for the formation of lamin-B mitotic spindle envelope and matrix. Interface Focus, 4:20130063. Doi: 10.1098/rsfs.2013.0063.
75. Chen H, Zheng X, and Zheng Y (2014). Age-Associated Loss of Lamin-B Leads to Systemic Inflammation and Gut Hyperplasia. Cell 159: 829–843.
76. Wan Y, Zheng, X, Chen H, Guo Y, Jiang H, He X, Zhu X, and Zheng Y (2015). Splicing function of mitotic regulators links R-loop mediated DNA damage to tumor cell killing. J. Cell Biol. 209: 235-246.
77. Zheng X, Kim Y, and Zheng Y (2015). Identification of lamin-B-regulated chromatin regions based on chromatin landscapes. Mol Biol Cell. 26:2685-2697.
78. Chen H, Zheng Y, and Zheng Y (2015). Lamin-B in systemic inflammation, tissue homeostasis, and aging. Nucleus 6:183-186.
79. Jiang H, He X, Feng D, Zhu X, and Zheng Y (2015). RanGTP aids anaphase entry through Ubr5-mediated protein turnover. J. Cell Biol. 211:7-18.
80. Oakley BR, Paolillo V, and Zheng Y (2015). Gamma tubulin complexes in microtubule nucleation and beyond. Mol. Biol. Cell. 26:2957-2962
81. Guo Y and Zheng Y (2015). Lamins position the nuclear pores and centrosomes by modulating dynein. Mol. Biol. Cell. 26:3379-3389.
82. Shimi T, Kittisopikul M, Tran M, Goldman AE, Adam SA, Zheng Y, Jaqaman K, Goldman RD (2015). Structural Organization of the Nuclear Lamin Isoforms in the Nuclear Lamina Revealed by Super Resolution Microscopy. Mol. Biol. Cell. 2015 Aug 26. pii: mbc.E15-07-0461. [Epub ahead of print].
83. Jiang H, Wang S, Huang Y, He X, Cui H, Zhu X, and Zheng Y (2015). Phase Transitions of Spindle-Associated Protein Regulate Spindle Apparatus Assembly. Cell 163:108-122.
84. Zheng X, Yue S, Chen H, Weber, B, Jia J, and Zheng Y (2015). Low cell number epigenome profiling aids the study of lens aging and hematopoiesis. Cell Report 13: 1-14.
85. Tran JT, Chen H, Zheng X, and Zheng Y (2016). Lamin in inflammation and aging. Invited review from Current Opinion in Cell Biology. Curr Opin Cell Biol. 40:124-130.
86. Chen H, Zheng X, Xiao D, and Zheng Y (2016). Age-associated de-repression of retrotransposons in the Drosophila fat body, the potential cause and consequences. Aging Cell, 3:542-52. doi: 10.1111/acel.12465. Epub 2016 Apr 12.
87. Tran J, Zheng X, and Zheng Y (2016). Lamin-B1 contributes to the proper timing of epicardial cell migration and function during embryonic heart development. Mol. Biol. Cell. 27:3956-3963. Epub 2016 Oct 19.
88. Zeituni EM, Wilson MH, Zheng X, Iglesias PA, Sepanski MA, Siddiqi MA, Anderson JL, Zheng Y, and Farber SA (2016). Endoplasmic reticulum lipid flux influences enterocyte nuclear morphology and lipid-dependent transcriptional response. J Biol Chem. 291:23804-23816.
89. Wall C, Dibattista M, Zheng X, Yue S, Young S, Reisert J, Zheng Y, and Zhao H (2017). Lamin-B1 is required for cell type-specific gene expression and terminal differentiation of olfactory sensory neurons. Nature Communication. Published online: doi: 10.1038/ncomms15098.
90. Zheng X, Zheng Y. CsoreTool: Fast Hi-C compartment analysis at high resolution. Bioinformatics. (2017) 34:1568-1570. Epub: doi: 10.1093/bioinformatics/btx802.
91. Huang Y, Li T, Ems-McClung SC, Walczak CE, Prigent C, Zhang X, and Zheng Y (2018). Aurora A activation in mitosis promoted by BuGZ. Journal of Cell Biology 217: 1077-116. Epub 2017 Oct 26th.
92. Zheng X, Hu J, Yue S, Kristiani L, Kim M, Kim Y,and Zheng Y (2018). Lamins organize the three dimensional genome from nuclear periphery in ES cells. Molecular Cell 71: 802-815.
93. Li B, Chen J, Chao, B, Zheng Y, and Xiao, X (2018). A lamin-binding ligand inhibits homologous recombination repair of DNA double stranded breaks. ACS Central Science 4:1201-1210.
94. Kim Y, Zheng X, and Zheng Y (2019). Role of lamins in 3D genome organization and global gene expression. Nucleus 10:33-41. 10.1080/19491034.2019.1578601.
95. Yue S, Zheng X, and Zheng Y (2019). Cell type-specific role of lamin-B1 and its inflammation-driven reduction in organ building and aging. Aging Cell https://dio.org/10.1111/acel.12952(link is external). (Aug;18(4):e12952. doi: 10.1111/acel.12952. Epub 2019 Apr 9.)
96. Liu W, Yue S, Zheng X, Cao J, Zheng Y (2018). aFARP-ChIP-seq, a convenient and reliable method for genome profiling in as few as 100 cells with a capability for multiplexing ChIP-seq. Epigenetics https://doi.org/10.1080/15592294.2019.1621139(link is external) (Jun 6:1-17. doi: 10.1080/15592294.2019.1621139. [Epub ahead of print]).
97. Ashish KT. and Zheng Y (2019). Protein Phase separation in mitosis. Current Opinion in Cell Biology 60:92-98.
98. Xiao Y, Chen J, Wan Y, Gao Q, Jing N, Zheng Y, and Zhu X (2019). Regulation of zebrafish dorsal ventral patterning by phase separation of RNA-binding protein Rbm14. Cell Discovery 5:37-54.
99. Hu Y, Zheng Y, Fan C-M, Zheng Y (2020). Single cell lineage dynamics of the endosymbiotic cell type in a soft coral Xenia species. Nature, 582 (7813): 534-538. DOI: 10.1038/s41586-020-2385-7.
100. Tran J. R, Paulson D. I., Moresco J. J., Adam S. A, Yates J. R. III3, Goldman R. D., and Zheng Y (2021). The versatility of Ascorbate Peroxidase-aided mapping uncovers insights of the nuclear lamina interactions and function. Journal of Cell Biology. Vol. 220: No. 1 e20202129.
101. Kittisopikul, M, Shimi, T., Tatli, M., Tran, J.R., Zheng, Y., Medalia, O., Jaqaman, K., Adam, S.A., Goldman, R.D., (2021). Nuclear Lamins A/C and B1 provide a structural framework that organizes and anchors Nuclear Pore Complexes. Journal of Cell Biology. Vol. 220 No. 4 e202007082.
102. Tran, Joseph R.; Zheng, Xiaobin; Adam, Stephen A.; Goldman, Robert D.; Zheng, Yixian (2022). High quality mapping of chromatin at or near the nuclear lamina from small numbers of cells reveals cell cycle and developmental changes of chromatin at the nuclear periphery. Nucleic acids research DOI: 10.1093/nar/gkac762.
103. Vahabikashi, Amir; Sivagurunathan, Suganya; Nicdao, Fiona Ann Sadsad; Han, Yu Long; Park, Chan Young; Kittisopikul, Mark; Wong, Xianrong; Tran, Joseph R.; Gundersen, Gregg G.; Reddy, Karen L.; Luxton, G. W. Gant; Guo, Ming; Fredberg, Jeffrey J.; Zheng, Yixian; Adam, Stephen A.; Goldman, Robert D. (2022). Nuclear lamin isoforms differentially contribute to LINC complex-dependent nucleocytoskeletal coupling and whole-cell mechanics. PROCEEDINGS OF THE NATIONAL ACADEMY OF SCIENCES OF THE UNITED STATES OF AMERICA. DOI: 10.1073/pnas.2121816119.
104. Chin, Alexander F.; Zheng, Yixian; Hilser, Vincent J. (2022). Phylogenetic convergence of phase separation and mitotic function in the disordered protein BuGZ. PROTEIN SCIENCE DOI: 10.1002/pro.4270.
105. Kono, Yohei; Adam, Stephen A; Sato, Yuko; Reddy, Karen L; Zheng, Yixian; Medalia, Ohad; Goldman, Robert D; Kimura, Hiroshi; Shimi, Takeshi. (2022). Nucleoplasmic lamin C rapidly accumulates at sites of nuclear envelope rupture with BAF and cGAS. The Journal of cell biology. DOI: 10.1083/jcb.202201024
106. Hu M, Bai Y, Zheng Z, and Zheng Y (2023). Coral-algal endosymbiosis characterized using RNAi and single-cell RNA-seq. Nature Microbiology, published online at: https://doi.org/10.1038/s41564-023-01397-9. Previously uploaded to bioRxiv. doi: https://www.biorxiv.org/content/10.1101/2022.12.13.520278v1. PMID: 37217718.
107. Zheng, Xiaobin; Joseph R. Tran; Zheng, Yixian. (2023) A model-based analysis reveals three dimensional genome organization heterogeneity and functional sub-compartments in cell populations. bioRxiv posting (2022): doi: https://doi.org/10.1101/2022.06.27.497761. Bioinformatics.
108. Yon WJ, Tran JR, Ha T, Zheng Y, and Pedersen RTA. BuGZ exhibits guanine nucleotide exchange factor activity toward tubulin. https://www.biorxiv.org/content/10.1101/2023.05.09.539990v2